Using EMLP in PyTorch¶
So maybe you haven’t yet realized that Jax is the best way of doing deep learning – that’s ok!
You can use EMLP and the equivariant linear layers in PyTorch. Simply replace import emlp.nn as nn with import emlp.nn.pytorch as nn.
If you’re using a GPU (which we recommend), you will want to set the environment variable so that Jax doesn’t steal all of the GPU memory from PyTorch. Note that if a GPU is visible under CUDA_VISIBLE_DEVICES, you must use the PyTorch EMLP on the GPU.
[1]:
%env XLA_PYTHON_CLIENT_PREALLOCATE=false
env: XLA_PYTHON_CLIENT_PREALLOCATE=false
[2]:
import torch
import emlp.nn.pytorch as nn
[3]:
from emlp.reps import T,V
from emlp.groups import SO13
repin= 4*V # Setup some example data representations
repout = V**0
G = SO13() # The lorentz group
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
[4]:
x = torch.randn(5,repin(G).size()).to(device) # generate some random data
model = nn.EMLP(repin,repout,G).to(device) # initialize the model
model(x)
[4]:
tensor([[-0.0039],
[-0.0041],
[-0.0042],
[-0.0040],
[-0.0039]], device='cuda:0', grad_fn=<AddmmBackward>)
The model is a standard pytorch module.
[5]:
model
[5]:
EMLP(
(network): Sequential(
(0): EMLPBlock(
(linear): Linear(in_features=16, out_features=419, bias=True)
(bilinear): BiLinear()
(nonlinearity): GatedNonlinearity()
)
(1): EMLPBlock(
(linear): Linear(in_features=384, out_features=419, bias=True)
(bilinear): BiLinear()
(nonlinearity): GatedNonlinearity()
)
(2): EMLPBlock(
(linear): Linear(in_features=384, out_features=419, bias=True)
(bilinear): BiLinear()
(nonlinearity): GatedNonlinearity()
)
(3): Linear(in_features=384, out_features=1, bias=True)
)
)
Example Training Loop¶
Ok what about training and autograd and all that? As you can see the training loop is very similar to the objax one in Constructing Equivariant Models.
[6]:
import torch
import emlp.nn.pytorch as nn
from emlp.groups import SO13
import numpy as np
from tqdm.auto import tqdm
from torch.utils.data import DataLoader
from emlp.datasets import ParticleInteraction
trainset = ParticleInteraction(300) # Initialize dataset with 1000 examples
testset = ParticleInteraction(1000)
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
BS=500
lr=3e-3
NUM_EPOCHS=500
model = nn.EMLP(trainset.rep_in,trainset.rep_out,group=SO13(),num_layers=3,ch=384).to(device)
optimizer = torch.optim.Adam(model.parameters(),lr=lr)
def loss(x, y):
yhat = model(x.to(device))
return ((yhat-y.to(device))**2).mean()
def train_op(x, y):
optimizer.zero_grad()
lossval = loss(x,y)
lossval.backward()
optimizer.step()
return lossval
trainloader = DataLoader(trainset,batch_size=BS,shuffle=True)
testloader = DataLoader(testset,batch_size=BS,shuffle=True)
test_losses = []
train_losses = []
for epoch in tqdm(range(NUM_EPOCHS)):
train_losses.append(np.mean([train_op(*mb).cpu().data.numpy() for mb in trainloader]))
if not epoch%10:
with torch.no_grad():
test_losses.append(np.mean([loss(*mb).cpu().data.numpy() for mb in testloader]))
Ok so it’s not nearly as fast as in Jax (maybe 15x slower), but hey you said you wanted PyTorch
[7]:
import matplotlib.pyplot as plt
plt.plot(np.arange(NUM_EPOCHS),train_losses,label='Train loss')
plt.plot(np.arange(0,NUM_EPOCHS,10),test_losses,label='Test loss')
plt.legend()
plt.yscale('log')
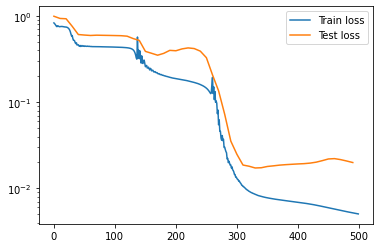
Bonus: Try out model=nn.MLP(trainset.rep_in,trainset.rep_out,group=SO13()).to(device) and see how well it performs on this problem.
Converting Jax functions to PyTorch functions (how it works)¶
You can use the underlying equivariant bases \(Q\in \mathbb{R}^{n\times r}\) and projection operators \(P = QQ^\top\) in pytorch also.
Since these objects are implicitly defined through LinearOperators, it is not as straightforward as simply calling torch.from_numpy(Q). However, there is a way to use these operators within PyTorch code while preserving any gradients of the operation. We provide the function emlp.reps.pytorch_support.torchify_fn to do this.
[ ]:
import jax
import jax.numpy as jnp
from emlp.reps import V
from emlp.groups import S
For example, let’s setup a representation \(S_4\) consisting of three vectors and one matrix.
[8]:
W =V(S(4))
rep = 3*W+W**2
First we compute the equivariant basis and equivariant projector linear operators, and then wrap them as functions.
[9]:
Q = (rep>>rep).equivariant_basis()
P = (rep>>rep).equivariant_projector()
[10]:
applyQ = lambda v: Q@v
applyP = lambda v: P@v
We can convert any pure pytorch function into a jax function by applying torchify_fn. Now instead of taking jax objects as inputs and outputing jax objects, these functions take in PyTorch objects and output PyTorch objects.
[11]:
from emlp.nn.pytorch import torchify_fn
applyQ_torch = torchify_fn(applyQ)
applyP_torch = torchify_fn(applyP)
As you would hope, gradients are correctly propagated whether you use the original Jax functions or the torchified pytorch functions.
[12]:
x_torch = torch.arange(Q.shape[-1]).float().cuda()
x_torch.requires_grad=True
x_jax = jnp.asarray(x_torch.cpu().data.numpy())
[13]:
Qx1 = applyQ(x_jax)
Qx2 = applyQ_torch(x_torch)
print("jax output: ",Qx1[:5])
print("torch output: ",Qx2[:5])
jax output: [0.48484263 0.07053992 0.07053989 0.07053995 1.6988853 ]
torch output: tensor([0.4848, 0.0705, 0.0705, 0.0705, 1.6989], device='cuda:0',
grad_fn=<SliceBackward>)
The outputs match, and note that the torch outputs will be on whichever is the default jax device. Similarly, the gradients of the two objects also match:
[14]:
torch.autograd.grad(Qx2.sum(),x_torch)[0][:5]
[14]:
tensor([-2.8704, 2.7858, -2.8704, 2.7858, -2.8704], device='cuda:0')
[15]:
jax.grad(lambda x: (Q@x).sum())(x_jax)[:5]
[15]:
DeviceArray([-2.8703732, 2.7858496, -2.8703732, 2.7858496, -2.8703732], dtype=float32)
So you can safely use these torchified functions within your model, and still compute the gradients correctly.
We use this torchify_fn on the projection operators to convert EMLP to pytorch.